Phylogenetics¶
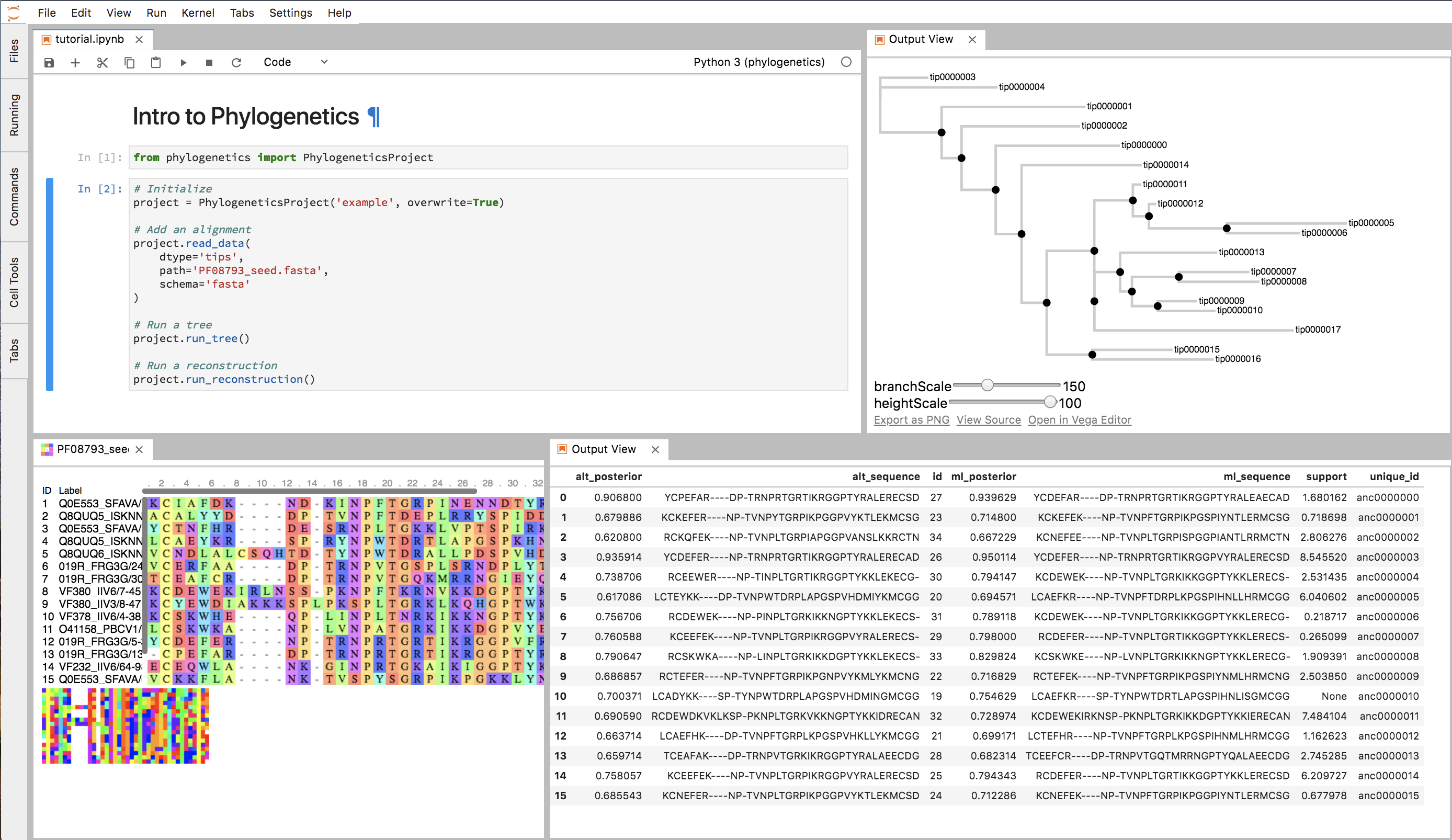
Phylogenetics is a minimal Python API for doing phylogenetics. It manages the annoying aspects of phylogenetics (i.e. file conversion) for you and lets you focus on exploring and interpreting the data.
Development goals¶
- docs:
- installation
- specific cases
- long branches
- what does branch support mean?
- sequence quality control
- alignment quality
- ancestors, posterior probabilities
- how do I come up with an answerable evolutionary question?
- design philosophy
- why do we need another package?
- as little low-level crap as possible (use Biopython, dendropy, etc.), let users interact simply via familiar csv and pandas df
- to implement:
- phylopandas (critical):
- uid/csv awareness
- tree/align integration (Zach)
- species correction
- datatype awareness (dna, rna, protein, codon)
- phylobot mk 2?
Installation¶
Install from PyPi:
pip install phylogenetics
To install a development version:
git clone https://github.com/Zsailer/phylogenetics
cd phylogenetics
pip install -e .
Dependencies¶
phylogenetics manages phylogenetics data. Currently, it doesn’t do any of the phylogenetic calculations itself. For this, use external tools like:
phylogenetics is built on top of following python stack:
- DendroPy: A Python library for phylogenetic scripting, simulation, data processing and manipulation.
- ToyTree: A minimalist tree plotting library using toyplot graphs
- PhyloPandas: Pandas for phylogenetics
- PyASR: Ancestral Sequence Reconstruction in Python